EHDS 2027 compliance deadline
The European Health Data Space Regulation mandates controlled secondary use of health data with full auditability and governance across member states by 2027.
EHDS · OMOP · Governed data sharing
Hydra helps healthcare organizations design, govern, anonymize, transform, and publish interoperable datasets for secondary use.
The platform combines HealthDCAT-AP catalogs, IDS/Gaia-X distribution patterns, OMOP modeling, and agent-assisted operations so data assets can move from inventory to controlled access.
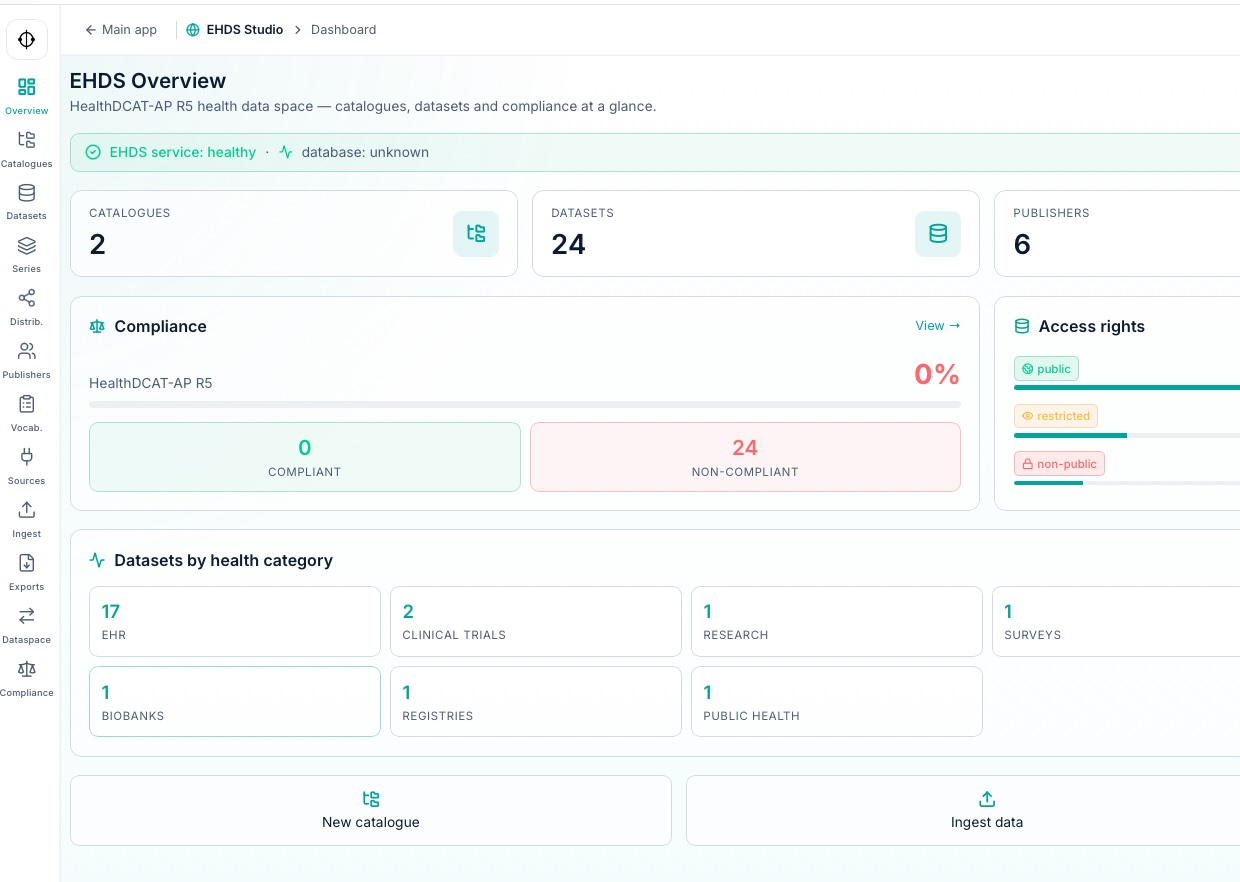
HealthDCAT-AP
metadata catalogs ready for European federation
8 connectors
TCP/MLLP, HTTP, files, databases, SMTP, WebSocket, and Kafka
18 PHI De-identification
automatic anonymization with auditability
OMOP CDM
research-grade modeling and vocabulary alignment
The European Health Data Space Regulation mandates controlled secondary use of health data with full auditability and governance across member states by 2027.
Patient records, cohort datasets, and research outputs are fragmented across hospitals, registries, and networks — inaccessible without a governed exchange layer.
Most teams apply de-identification and cataloging by hand — no standard format, no audit trail, and no alignment with HealthDCAT-AP or OMOP for federated access.
Federated cohort studies, real-world evidence, and registry queries require vocabulary-aligned, OMOP-ready datasets — without a common model, cross-institutional research stays manual and error-prone.
Platform architecture
A single platform covers the full journey: catalog, ingestion, anonymization, OMOP modeling, and governed distribution to European health data infrastructure.
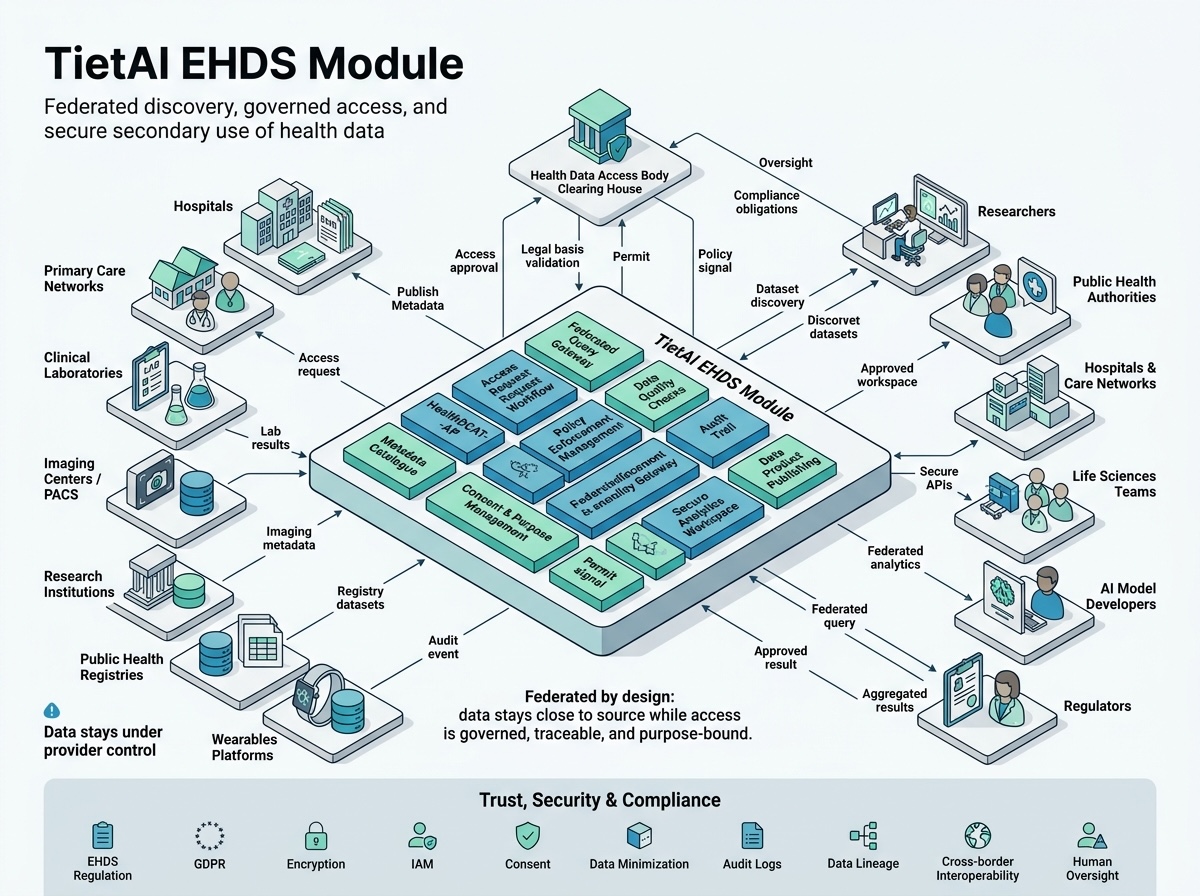
The EHDS layer focuses on cataloging, interoperability, anonymization, distribution, and policy-aware exchange across healthcare and research networks.
Create standardized dataset and data service metadata, including RDF export for future European health data federation.
Connect FHIR, OLTP/OLAP, files, databases, HTTP, TCP/MLLP, WebSocket, Kafka, SMTP, and other operational systems.
Detect and treat identifiers in structured and unstructured data with redaction, markers, masking, encryption, and synthetic replacement.
Publish downloadable distributions or API-based access across multiple destinations, formats, and health data categories.
Define assets, policies, contracts, negotiation flows, and transfer monitoring for interoperable data space operations.
Accelerate catalog creation, asset preparation, engineering workflows, and operational publication with guided agents.
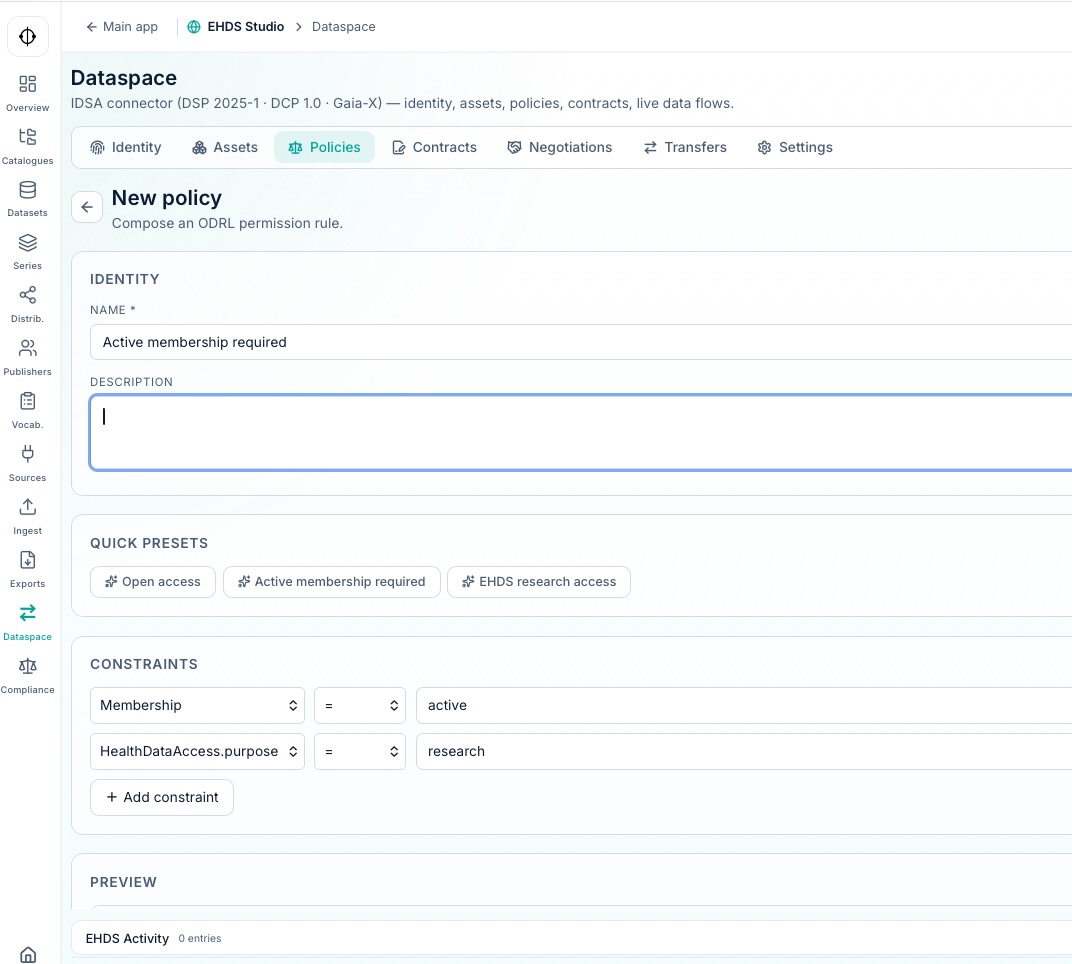
Rules and governance
Hydra generates a complete governance layer across every dataset in the data space, following TEHDAS2 recommendations for policy definition, dataset lifecycle control, auditability, and controlled secondary use.
This ruleset can be published through a range of connectors compliant with the DSP protocol and IDSA recommendations. Hydra also implements the National Contact Point (NCP) architecture described in TEHDAS2 M7.3, based on a two-layer model: Business Layer (Dispatcher) and Messaging Layer (eDelivery/Domibus).
The OMOP layer supports modeling, vocabulary validation, documentation, export, and knowledge-driven enrichment for research and secondary use.
Map source datasets into the OMOP Common Data Model using repeatable ingestion, transformation, and validation workflows.
Validate ICD-10, ICD-11, SNOMED CT, LOINC, ATC, ICPC-2, HPO, OMOP CDM, and UMLS-linked concepts.
Generate checks, transformation documentation, and controlled exports so datasets remain traceable and reusable.
Use natural language processing models to detect clinical entities in text and feed structured modeling pipelines.
Use graph-based terminology services, ontology navigation, semantic search, and agent-friendly APIs for concept alignment.
Produce interoperable data services and dataset distributions for research, analytics, AI agents, and regulated exchange.
Start with catalog management and data inventory, move into ingestion and transformation, then publish controlled distributions for download or API consumption.
Governance, anonymization, vocabulary validation, and policy handling stay attached to the dataset lifecycle instead of becoming manual side work.
Build the catalog, governance, OMOP modeling, and distribution layer needed for secondary use and European health data collaboration.